ChimeRMarker® Automated Chimerism Analysis Software
ChimeRMarker® automated chimerism analysis software integrates speed and accuracy with a biologist-friendly interface. The software can be used to monitor chimerism level in both an allogeneic stem cells transplant (SCT) or a hematopoietic stem cells transplant (HSCT), bone marrow transplant (BMT, post bone marrow engraftment), and cord and peripheral blood stem cells transplant (PBSCT) samples.
Complete Chimerism Detection, Long-term Monitoring, and Quantification in one program!
Increase analysis capacity 20-30% annually (Read the Association of Molecular Pathology presentation)
ChimeRMarker software provides:
- Accurate, rapid genotyping and chimerism analysis with documented time savings of up to 85%
- Automatically identifies donor and recipient peaks in post-BMT samples
- Calculates percent chimerism and quality metrics for single donor or double donor cases
- Easily appends for longitudinal monitoring post-BMT
- Multi-lineage capabilities for chimerism analysis of T-cells, B-cells, and other cell type populations
ChimeRMarker software includes functions for comparison of samples at different time points to conduct longitudinal studies for monitoring each individual and a comprehensive chimerism analysis report. The chimerism analysis performs repetitive calculations (using published methods. Dr. Don Kristt).
ChimeRMarker software also has a linked maternal cell contamination (MCC) application. ChimeRMarker software is compatible with ABI®PRISM, Applied Biosystems® SeqStudio, Hiachi High-Tech Compact CE Sequencer DS3000, Promega Spectrum Compact CE Systems and Syntol Nanophore® 05 genetic analyzers, and custom primers or commercially available human identification chemistries for STR genotyping (including, but not limited to Identifiler®, GlobalFiler™ 6-Dye™, PowerPlex®16, PowerPlex®ESI, GenePrint® 24, PowerPlex®Fusion 5 and 6 Dye, Investigator®24Plex). Chimerism analysis is completely integrated to the main analysis screen, removing the error-prone step of data transfer from genotyping software to calculation spread sheets.
Click here to read “Analytic Approach for Complete and Mixed Engraftment Analysis using Commercial Software”, presented at Applied Molecular Pathology meetings 2016 by Lea F Surrey MD, Jennifer R Smith BA, Daniel Lubin MD, Vivianna M. Van Deerlin MD PhD, and Christopher D Watt MD PhD.
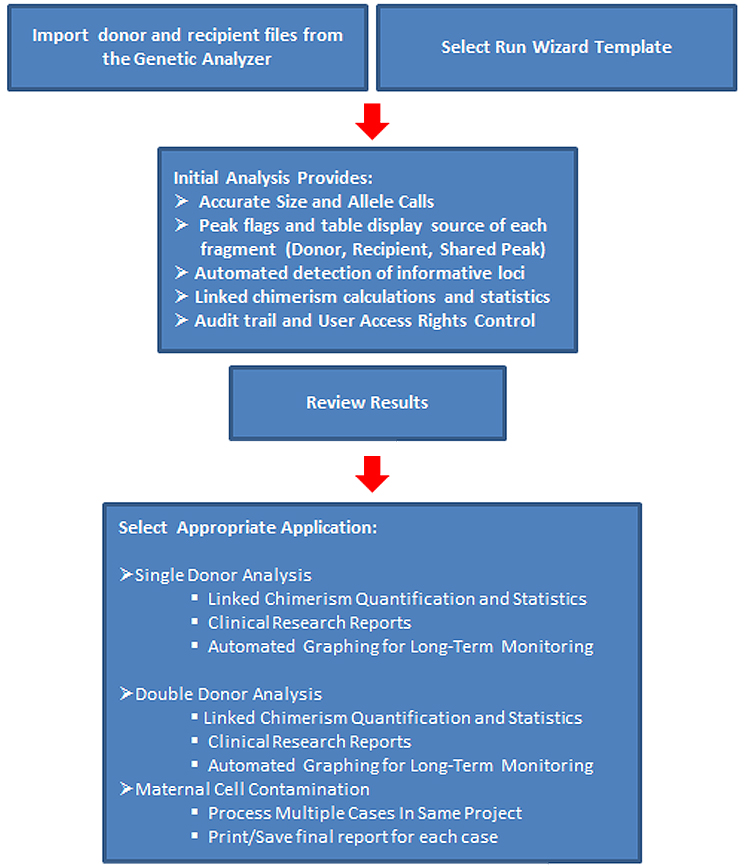
*Time study reported by Franciscan Alliance, St. Francis Health, Indianapolis, IN
Application Notes:
- ChimeRMarker® Software for Automated Analysis, Chimerism Detection, Quantification and Monitoring from Short Tandem Repeat (STR) DNA in Post-Transplant Samples App Note
- Automated Chimerism Analysis Using ChimeRMarker® Software: A Concordance Study
Helpful Links:













